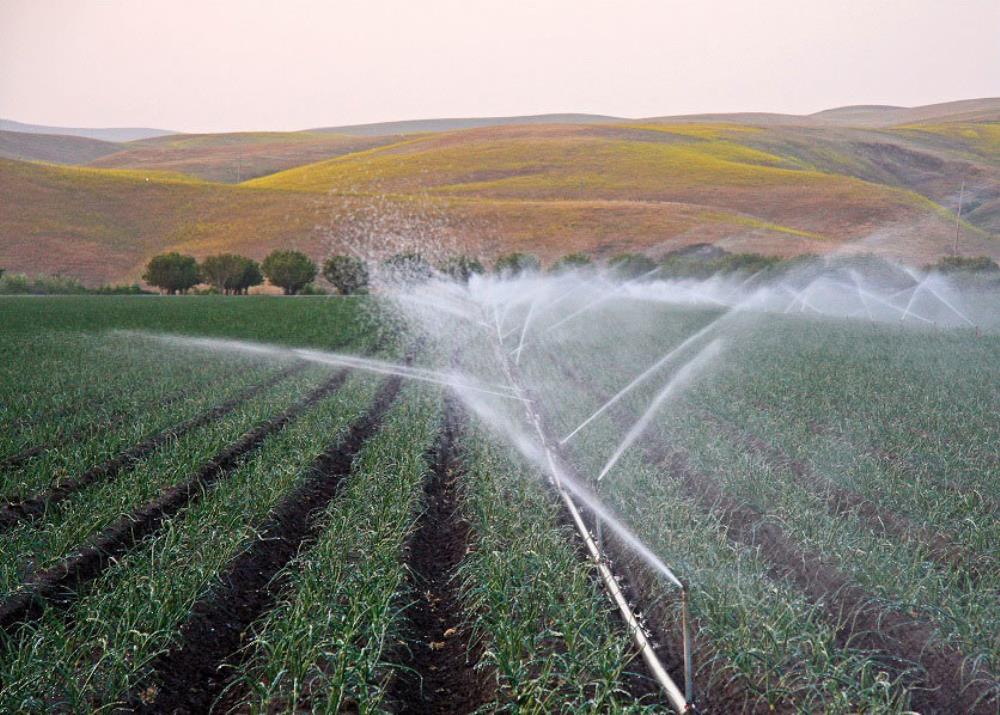



WaterNet
for
pour le
pour le
Canada
Menu
 WaterNet
WaterNetHome
 GWFO
GWFOHome
 Master
MasterList
 Data
DataCentre
 Collections
Collections
X
Defaults
Select All
 Websites
Websites
X
Global Water Futures Observatories (GWFO)
Global Water Futures (GWF)
Global Institute for Water Security (GIWS)
International Network of Alpine Research Catchment Hydrology
 Legacy sites
Legacy sites
Legacy Research Programs
X
Changing Cold Regions Network (CCRN)
Drought Research Initiative (DRI)
International Network of Alpine Research Catchment Hydrology (Legacy Site)
Improving Processes & Parameterization for Prediction in Cold Regions Hydrology (IP3)
The Mackenzie Global Energy and Water Cycle Experiment (GEWEX) Study (MAGS)
 Legacy sites
Legacy sites
 Map
Map
 Utilities
Utilities
 . . .
. . .
X
Clear
Select All
Advanced Search
 Metadata Editor
Metadata Editor
 Alias List Editor
Alias List Editor